AmberMD Agent
A fully autonomous molecular dynamics agent powered by Claude Code.
Describe what you want to simulate — in plain language — and the agent handles everything else.
It searches six live science databases to gather structures, ligand parameters, and experimental
binding data, reads the Amber manual to plan the correct protocol, writes all input files,
submits jobs to your HPC cluster via SLURM, monitors progress, and interprets the results.
The Amber manual is bundled with the code and indexed for instant lookup — no manual setup required.
Whether it's a standard MD simulation, thermodynamic integration, umbrella sampling, MM-GBSA,
or replica exchange — just describe the system and the question you want to answer.
The agent reads the manual, builds the workflow from scratch, and executes it end to end.
Workflow
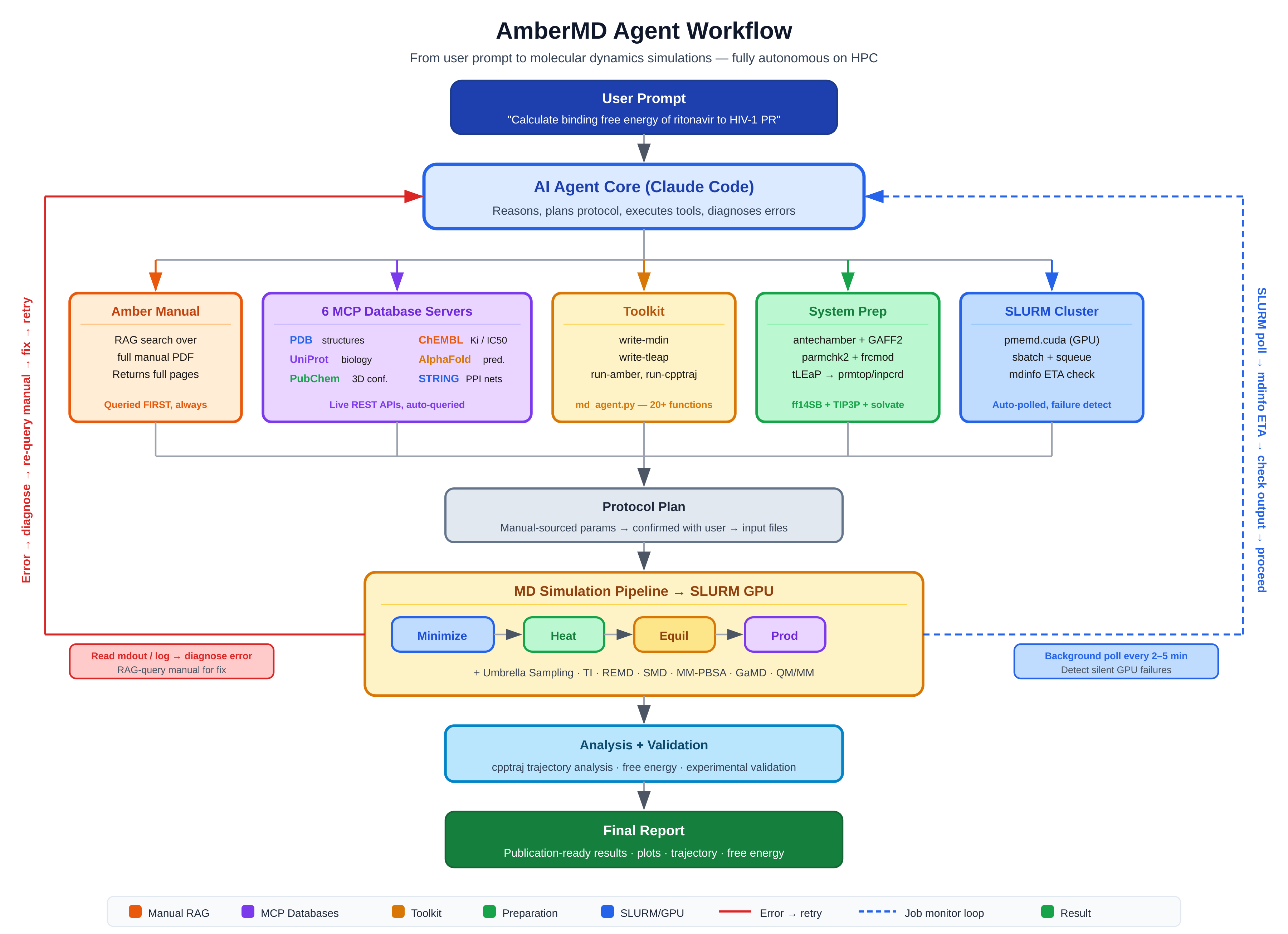
How It Works
┌─────────────────────────────────────────────────────────────┐
│ User: "Calculate binding free energy of erlotinib to EGFR" │
│ │
│ Agent: │
│ 1. Queries live databases via MCP │
│ PDB → find best EGFR structure + validate quality │
│ UniProt → kinase domain boundaries, known mutations │
│ PubChem → erlotinib SMILES, 3D conformer for params │
│ ChEMBL → experimental Ki to validate results against │
│ │
│ 2. Queries Amber manual via RAG │
│ → "thermodynamic integration setup" │
│ → "softcore potentials ifsc" │
│ → "lambda schedule recommendations" │
│ │
│ 3. Plans: "Based on the manual, I'll use TI with..." │
│ - 11 lambda windows (0.0 → 1.0) │
│ - GAFF2 for erlotinib via antechamber │
│ - ff14SB + TIP3P │
│ - SLURM array job, one GPU per window │
│ │
│ 4. Executes using toolkit functions │
│ write_mdin() → run_amber() → sbatch → analyze │
│ │
│ 5. Validates: computed ΔG vs ChEMBL experimental Ki │
└─────────────────────────────────────────────────────────────┘
The AI is the brain. The toolkit is the hands. The manual is the textbook. MCP servers are the library.
Architecture
amber-md-agent/
├── md_agent.py # Toolkit: PDB ops, file writers, runners, RAG, SLURM
├── CLAUDE.md # Agent instructions (protocol, rules, known issues)
├── scripts/
│ ├── slurm_template.sh # One-time cluster config — edit once, used everywhere
│ └── cap_protein.py # ACE/NME terminal capping utility
├── mcp_servers/ # Live database integrations (6 servers)
│ ├── pdb_server.py # RCSB PDB: structure search, validation, ligand info
│ ├── uniprot_server.py # UniProt: protein info, domains, mutations, PTMs
│ ├── pubchem_server.py # PubChem: compound search, 3D conformers, bioassays
│ ├── alphafold_server.py # AlphaFold DB: predicted structures, pLDDT confidence
│ └── stringdb_server.py # STRING-DB: PPI networks, pathway enrichment
├── references/
│ └── (place Amber manual PDF here → gets indexed for RAG)
└── studies/ # All simulation work lives here
└── <study_name>/
├── raw_pdbs/ # Downloaded PDB files
├── system/ # Topology prep (tleap, prmtop, inpcrd)
├── simulations/ # min1/ min2/ heat/ equil/ prod/
├── analysis/ # cpptraj, .dat files, plots/
└── logs/
... [View full README on GitHub](https://github.com/nagarh/amber-md-agent#readme)
